Å@3. ER-asociated degradation of misfolded proteins by EDEM-ERdj5 system
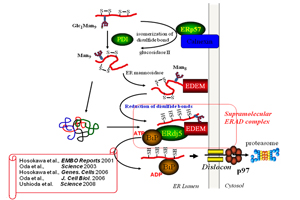 Å@We are working on the molecular mechanism of protein quality control by oxidoreductases in the ER. We identified that an ER disulfide reductase, ERdj5 promotes ERAD by cleaving of intermolecular disulfide bonds of misfolded proteins and preventing their oligomer or aggregate formation. ERdj5 interacts with EDEM, which recognizes misfolded proteins via their N-glycan portions, and a molecular chaperone, BiP. This ternary complex composing with ERdj5, EDEM and BiP accelerates the ERAD. After domain analysis of ERdj5, we found that EDEM interacts with ERdj5 via the C-terminal thioredoxin-like domain cluster and recruits misfolded glycoproteins to ERdj5 and thioredoxin-like domains at the C-terminal cluster are involved in cleaving the intermolecular disulfide bonds of misfolded proteins. BiP associates with ERdj5 through the J-domain in the N-terminus, and captures the reduced misfolded proteins until they are transferred to the dislocation channel.
Å@We are working on the molecular mechanism of protein quality control by oxidoreductases in the ER. We identified that an ER disulfide reductase, ERdj5 promotes ERAD by cleaving of intermolecular disulfide bonds of misfolded proteins and preventing their oligomer or aggregate formation. ERdj5 interacts with EDEM, which recognizes misfolded proteins via their N-glycan portions, and a molecular chaperone, BiP. This ternary complex composing with ERdj5, EDEM and BiP accelerates the ERAD. After domain analysis of ERdj5, we found that EDEM interacts with ERdj5 via the C-terminal thioredoxin-like domain cluster and recruits misfolded glycoproteins to ERdj5 and thioredoxin-like domains at the C-terminal cluster are involved in cleaving the intermolecular disulfide bonds of misfolded proteins. BiP associates with ERdj5 through the J-domain in the N-terminus, and captures the reduced misfolded proteins until they are transferred to the dislocation channel.
Å@We have also established the interaction networks among around 20 ER oxidoreductases by co-immunoprecipitation and following LC/MS analysis to elucidate redox cascades in the ER. Based on the analysis, we have found that the redox complex composed of an ER oxidase, Ero1L and an ER oxidoreductase, PDI makes a regulatory hub complex, which oxidizes other ER oxidoreductases efficiently. We have also found that an ER oxidoreductase, TMX4, which is localized on the ER membrane, interacts with a lectin-like chaperone, calnexin and an ER oxidoreductase, ERp57, both of which are involved in the glycoprotein oxidative folding processes.
|
 TOP
TOP
 TOP
TOP